Centro di Ateneo per il Calcolo nella Scienza e nella Tecnica (CAST)
Interaction site of Epigallocatechin-3-gallate with Insulin aggregates
Risultato di uno sforzo di sperimentazione rigorosa, di un costante sviluppo di nuove tecnologie e di una continua attività teorica sono quelle che, in ogni dato periodo storico, chiamiamo le leggi fondamentali della natura. Nel corso del tempo nuovi e più precisi risultati sperimentali, una più attenta e illuminata sintesi teorica modificano continuamente queste leggi mirando a renderle sempre più fondamentali e più generali, capaci cioè di descrivere il maggior numero possibile di fenomeni in termini del minor numero possibile di assunzioni e di costituenti elementari.
La crescente compattezza delle teorie rende di contro sempre più difficile derivare da esse la spiegazione quantitativa di fenomeni legati a sistemi composti da molte parti in reciproca interazione, quali ad esempio sistemi percepibili su scala umana.
L’interesse nei sistemi macroscopici nasce da diverse esigenze. Da un punto di vista prettamente scientifico, una teoria può ritenersi “giusta” solo se regge il confronto con l’esperimento entro la precisione dello stesso; se tale confronto non è possibile a causa della difficoltà di derivare le predizioni teoriche non si può scientificamente asserire di avere compreso il sistema in esame. Dal un punto di vista tecnologico, gli oggetti di uso comune e la materia vivente rappresentano esempi concreti di sistemi molto complessi che incidono nella nostra vita quotidiana.
Previsioni sulla fenomenologia di sistemi anche estremamente complessi si possono ottenere in pratica ricorrendo al calcolo numerico: si considera un numero grande ma finito di costituenti elementari e si utilizza un calcolatore numerico per la soluzione delle equazioni matematiche che governano le interazioni di questi ultimi. Le equazioni differenziali vengono “discretizzate” e l'evoluzione del sistema viene riprodotta per un certo numero di tempi campionati ad intervalli piccoli ma finiti. Il grado di complessità dei problemi risolubili risulta pertanto essere correlato alla potenza di calcolo a disposizione e all’efficienza degli algoritmi numerici utilizzati.
Il CAST e' membro del nodo IT-SIMUL del CECAM insieme a La Sapienza, Politecnico di Torino, Politecnico di Milano, Universita' dell'Aquila, e IIT-Genova.
Obiettivi del CAST
Lo scopo del centro CAST negli ultimi anni è stato quello di rispondere alle esigenze prioritarie della comunità scientifica e teconologica del nostro Ateneo che si occupa della soluzione di problemi complessi.
Il CAST si è proposto e si propone per il futuro di fornire le necessarie risorse di calcolo alla comunità scientifico-teconologica del nostro Ateneo ma anche, e principalmente, di creare un linguaggio comune e una collaborazione scientifica tra i diversi gruppi che hanno avuto e avranno accesso a tali risorse di calcolo.
Tali obiettivi per gruppi scientifico-teconologici che si occupano di problemi derivanti dallo studio della biologia molecolare, dell'astrofisica, della fenomenologia delle particelle elementari, della meteorologia, della turbolenza, della fisica e chimica dei materiali etc. sono realizzabili sotto la spinta della comune esigenza di migliorare costantemente gli algoritmi numerici, le tecniche di programmazione, la teoria stessa dei problemi a molti corpi e del calcolo numerico e tutte le ulteriori problematiche ad esse correlate, aspetti tutti sviluppabili e condivisibili in modo interdisciplinare.
Contatti:
coordinatore: Prof. Olivia Pulci
Alcuni esempi di attivita' entro il CAST sono qui sotto elencati:
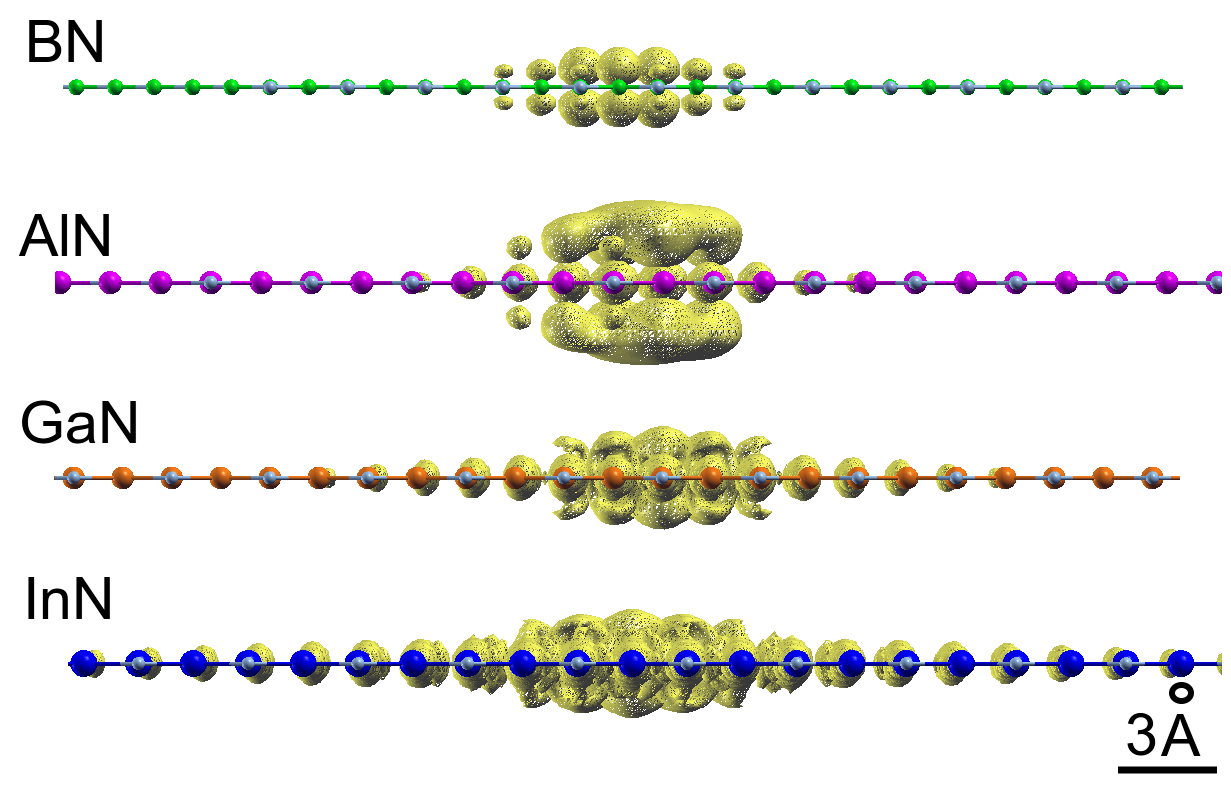
Excitons wavefunctions in 2D Nitrides
New Materials
The main research lines of the group of Theoretical Physics of Condensed Matter
are dedicated to:
i) investigate with theoretical and computational quantum mechanical methods the structural, electronic and optical properties of materials
ii) unveal the role of the effects of electronic correlation at equilibrium and out of equilibrium
iii) to develop theoretical-computational methods for the description of both steady-state and time-resolved ultra-fast electronic and optical spectroscopies
Keywords: Low-dimensional excitons, Opto-electronics and energy storage, Topological Properties of Matters, Correlation effects in organic materials
Highlights:
[1] 2N+ 4-rule and an atlas of bulk optical resonances of zigzag graphene nanoribbons, R. B. Payod et al., Nat. Comm. 11, 1-9, (2020)
[2] Optical Conductivity of Two-Dimensional Silicon: Evidence of Dirac Electrodynamics, C. Grazianetti et al., Nano Letters 18(11), pp. 7124-7132
(2018)
[3] "Theory and ab initio computation of the anisotropic light emission in monolayer transition metal dichalcogenides" HY Chen, M Palummo, D Sangalli, M Bernardi, Nano letters 18 (6), 3839-3843 (2018)
Contacts:
Prof. Olivia Pulci
Prof. Gianluca Stefanucci
Prof. Maurizia Palummo
Dr. Enrico Perfetto
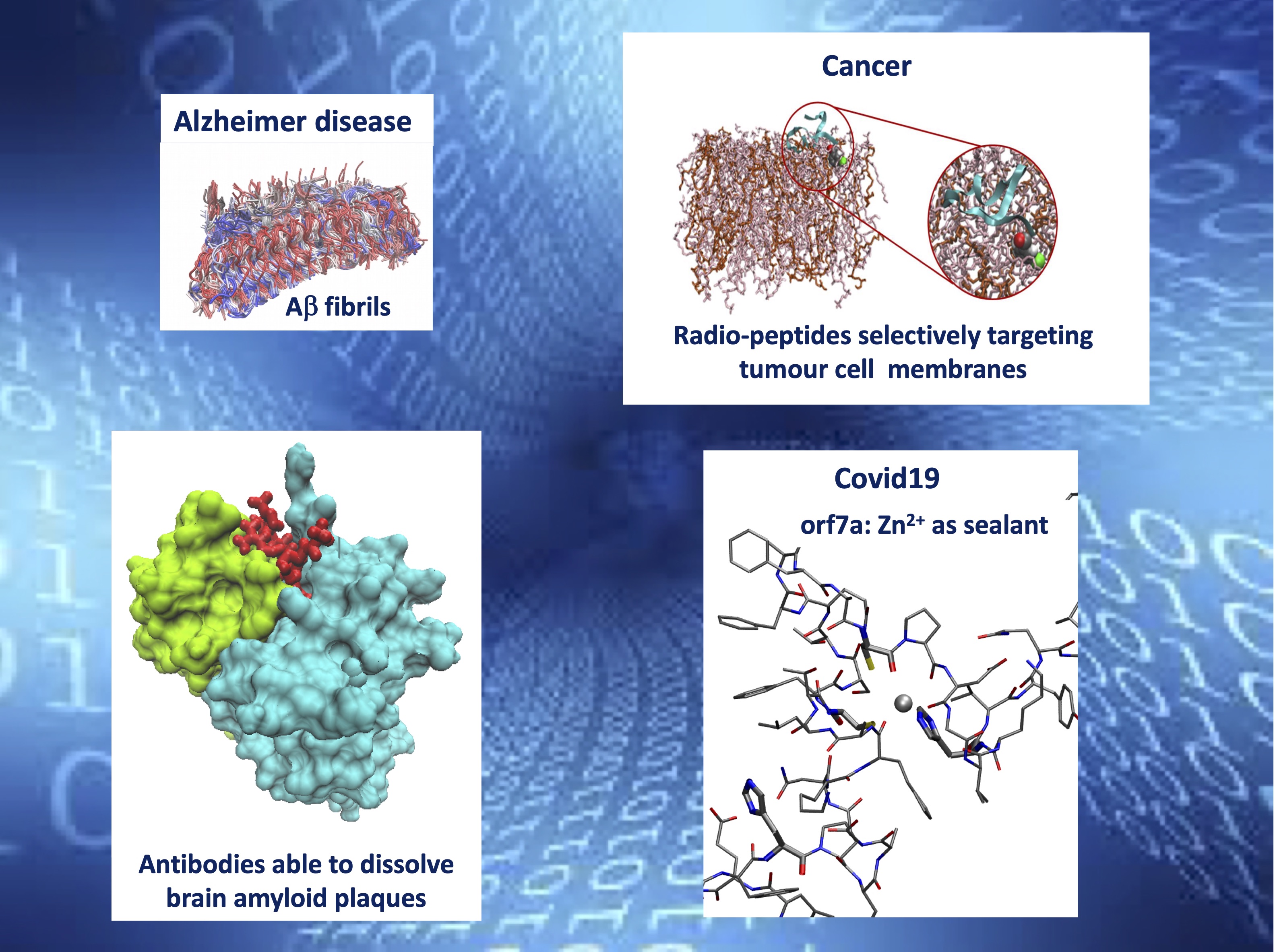
Biophysics:
There exist a large number (more than 50) of diseases associated to misfolding followed by aggregation of some specific protein. They are collectively called Protein Conformation Diseases (PCD’s). It is remarkable that all the known neurodegenerative diseases belong to the PCD class. In the effort of explaining how misfolding may give rise to PCD, computational approaches have proved to be on unreplaceable tool. Understanding mis-folding is a challenging problem that requires the systems of interest (aggregated proteins in solution, comprising 106 atoms) be studied at different time scales (from picosec to millisec) with the appropriate numerical approach. Within our research group there is also a strong experimental activity aimed at understanding the role of metal ions in mis-folding and aggregation. In this context simulations are exploited to build structural models to correctly interpret experimental results.
Key-words: Protein Conformation Diseases (PCD’s); Multi-level simulations; proteins sealed by metal ions.
Highlights
[1] V. Minicozzi, F. Stellato, M. Comai, M. Dalla Serra, C. Potrich, W. Meyer-Klaucke, S. Morante. “Identifying the Minimal Cu and Zn Binding Site Sequence in Amyloid Beta Peptides” J. Biol. Chem.283: 10784-10792 (2008).
[2] P. Giannozzi, K. Jansen, G. La Penna, V. Minicozzi, S. Morante, G.C. Rossi, F. Stellato “Zn induced structural aggregation patterns of -amyloid peptides by first-principle simulations and XAS measurements” Metallomics. 4(2):156-165 (2012).
[3] V. Minicozzi, R. Chiaraluce, V. Consalvi, C. Giordano, C. Narcisi, P. Punzi, G.C. Rossi and S. Morante, “Computational and experimental casus on beta sheet breakers targeting A1-40 fibrils.” J. Biol. Chem. (2014) 289,11242-11252.
[4] F. Stellato, M. Calandra, F. D’Acapito, E. De Santis, G. La Penna, G. Rossi, and S. Morante, “Multi-scale theoretical approach to X-ray absorption spectra in disordered systems: an application to the study of Zn(II) in water” (2018) PCCP 20 (38), 24775-24782.
Contacts:
Prof. Silvia Morante
Dr. Velia Minicozzi
Dr. Francesco Stellato
Molecular Biology:
1) Experimental and simulative characterization of DNA nanocages for a targeted
drug delivery.
2) Role of gut microbiota on Parkinson disease through metagenomics and machine learning approaches.
3) Identification of new compounds poisoning human topoisomerase 1 through experiments and molecular modeling.
Keywords MD simulations- Molecular modeling-metagenomics- DNA nanostructures
Highlights
[1] Raniolo S, Vindigni G, Unida V, Ottaviani A, Romano E, Desideri A, Biocca S. Entry, fate and degradation of DNA nanocages in mammalian cells: a matter of receptors. Nanoscale. 2018 Jul 5;10(25):12078-12086.
[2] Pietrucci D, Cerroni R, Unida V, Farcomeni A, Pierantozzi M, Mercuri NB, Biocca S, Stefani A, Desideri A. "Dysbiosis of gut microbiota in a selected population of Parkinson's patients." Parkinsonism Relat Disord. 2019 Aug;65:124-130.
[3] Oliveira CG, Romero-Canelón I, Silva MM, Coverdale JPC, Maia PIS, Batista AA, Castelli S, Desideri A, Sadler PJ, Deflon VM. "Palladium(ii) complexes with thiosemicarbazones derived from pyrene as topoisomerase IB inhibitors." Dalton Trans. 2019 Nov 12;48(44):16509-16517.
Contacts:
Prof. Alessandro Desideri desideri@uniroma2.it
Theoretical Particle Physics
Our research activity is mostly focused on the theory and phenomenology of strong interacting particles that we investigate by using Non-Perturbative Lattice techniques. We search for signals of new physics by performing precision
tests of the Standard Model of fundamental interactions. To this end, recently, we have developed state-of-the-art theoretical methods and computer codes to simulate numerically strong and electromagnetic interactions and computed, from the very first principles, fundamental quantities such as the masses of the nucleons, the leptonic decay rates of pseudoscalar mesons, etc. From these
calculations we extracted, at an unprecedented level of accuracy, fundamental parameters of the theory such as quark masses and CKM matrix elements.
key words: Theoretical Particle Physics, Lattice QCD+QED, Precision tests of the SM, CKM matrix elements.
Highlights:
[1] D.Giusti, V.Lubicz, G.Martinelli, C.Sachrajda, F.Sanfilippo, S.Simula, N.Tantalo
and C.Tarantino, ``First lattice calculation of the QED corrections to leptonic decay rates,'' Phys. Rev. Lett. 120 (2018) 7, 072001 doi:10.1103/PhysRevLett.120.072001
[arXiv:1711.06537 [hep-lat]].
[2] B.Lucini, A.Patella, A.Ramos and N.Tantalo,
``Charged hadrons in local finite-volume QED+QCD with C^⋆ boundary
conditions,'' JHEP 02 (2016), 076 doi:10.1007/JHEP02(2016)076 [arXiv:1509.01636 [hep-th]].
[3] de Divitiis, G.M. et al., [RM123 Collaboration],``Leading isospin breaking effects on the lattice, '' Phys. Rev. D 87 (2013) 11, 114505 doi:10.1103/PhysRevD.87.114505 [arXiv:1303.4896 [hep-lat]].
Contacts:
Prof. Nazario Tantalo
Dr. Giulia de Divitiis
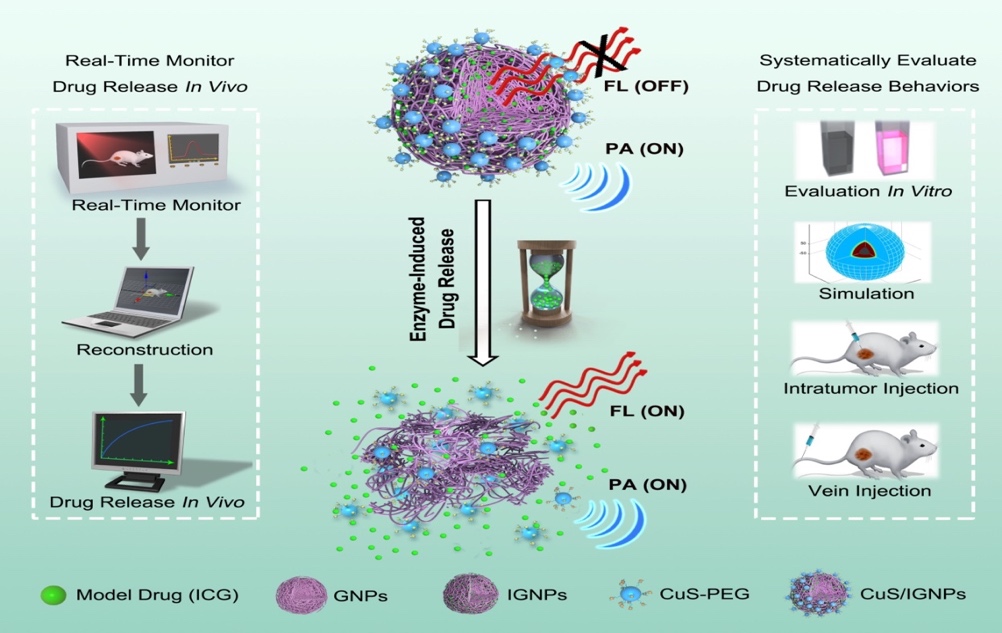
Development of an enzyme-activated anti-cancer approach.
Biophysics and Biomedical Engineering
Nicola Rosato, PhD in Biophysics (Pisa), is Professor of Biochemistry, at the University of Rome-Tor Vergata and Director of Clinical Engineering Service of Tor Vergata University Hospital. His scientific interest (H-index 34; Citations 3400) focuses upon structure and functions of proteins investigated by spectroscopical techniques, nanomedicine and nano biomaterials and development
of new medical devices.
Key words: fluorescence spectroscopy, nanomedicine, nano biomaterials, medical
devices
Highlights:
[1] Bottini M, Sacchetti C, et al. Targeted nanodrugs for cancer therapy: prospects and challenges. J Nanosci Nanotechnol. 2014; 14(1): 98-114.
[2] Motevalli S, Eltahan AS, et al. Co-encapsulation of curcumin and doxorubicin in albumin nanoparticles blocks the adaptive treatment tolerance of cancer cells. Biophysics Reports. 2019; 5: 19-30.
[3] Li X, Bottini M, et al. Core-satellite nanomedicines for in vivo real-time monitoring of enzyme-activatable drug release by fluorescence and photoacoustic dual-modal imaging. ACS Nano. 2019; 13(1): 176-186.
Prof. Nicola Rosato
Contact: nicola.rosato@uniroma2.
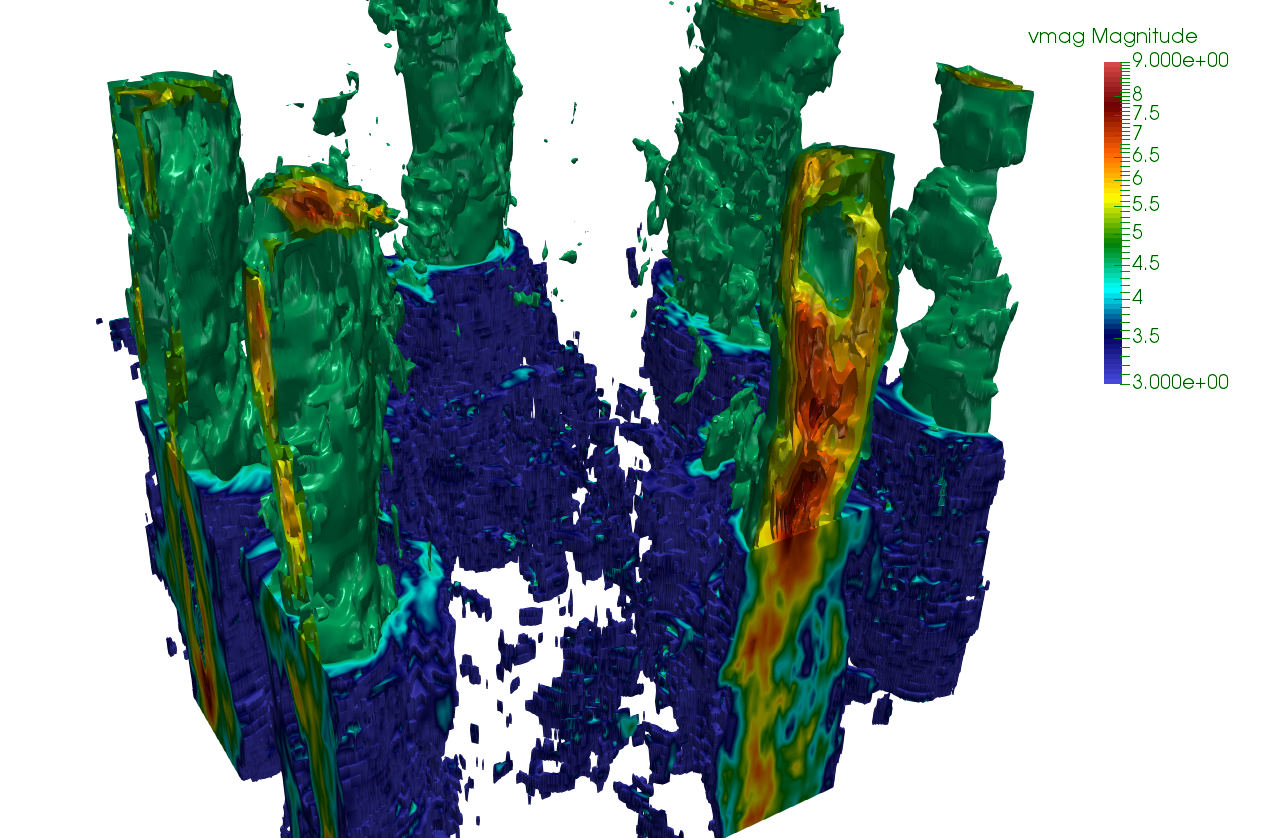

Fully developed rotating turbulent flows. The picture represents the velcoity intensity concentrated on the highly choerent cyclonic vortex structures
Complex fluids
The group working on complex flows and complex fluids in the Department of Physics is involved in high performance computing of fully developed turbulence, both Eulerian and Lagrangian, including applications to rotating and stratified
fluids, thermal convection and other bounded flows. The activity involving micro- and nano-scale flows is focused on non-Newtonian effects induced by confinement on foams and emulsions, including the complex physics of thermally
activated multi-phase flows. Tools used go from direct numerical simulations based on pseudospectral code, Lattice Boltzmann algorithms or big-data approaches based on Machine Learning and Reinforcement Learning. Recently, also
an activity on complex networks has started. People involved: Prof. R. Benzi, Prof. L. Biferale, Prof. M. Sbragaglia, Dott. M. Buzzicotti, Dott. G. Cimini as well as a number of post-docs and Phd students.
Keywords: large scale turbulence, micro- and nano-fluidics, foams, emulsions, Machine Learning for fluid dynamics, complex systems, High Performance Computing.
Highlights:
[1] Zermelo's problem: Optimal point-to-point navigation in 2D turbulent flows
using Reinforcement Learning
L Biferale, F Bonaccorso, M Buzzicotti, PC Di Leoni, K Gustavsson
Chaos 29, 103138, 2019
[2] Patricio Clark Di Leoni, Andrea Mazzino, and Luca Biferale Phys. Rev. X 10,
011023. 2020--
[3] Unified Theoretical and Experimental view on Transient Shear banding
by Benzi R., Divoux T., Barentin C., Manneville S., Sbragaglia M. and
Toschi F. Phys. Rev. Lett. 123, 248001 (2019)
Contacts:
Prof. Roberto Benzi
Prof. Luca Biferale
Prof. Mauro Sbragaglia
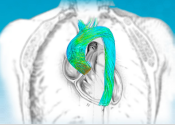
Digital twin of the aortic arch defined to support medical doctors in aneurysms prevention and correction
The Medical Digital Twin
Our research on the medical digital twin (DT) aims to develop state-of-the-art image based medical DTs for cardiovascular and orthopaedic applications based on patient specific data. DT applications include prevention, treatment and surgical planning. The research covers five research tracks
- high fidelity CAE multi-physics simulation with RBF mesh morphing (FEM, CFD, FSI, inverse FEM),
- Real time interaction with the DT by Augmented Reality, Haptic Devices and ROM,
- HPC tools, including GPUs, and cloud-based paradigms for fast and automated CAE processing of clinical database,
- Big Data management for population of patients imaging data and high fidelity CAE DTs,
- Additive Manufacturing of physical mock-up for surgical planning and training
that are mixed to gain a comprehensive medicine 4.0 approach toward an effective and innovative workflow. Our project is driven by the multi-disciplinary and multi-sectoral needs of a research network that counts more than 20 international partners (clinical, academic and industrial) that are active players of a strategic sector of the European medical and simulation industry (clinical experts, engineering analysts and simulation software technology developers). Learn more about cardiovascular applications on the MeDiTATe web page (meditate-project.eu).
Keywords
Computer Aided Engineering, Computational Fluid dynamics, Radial Basis Functions, Digital Twins, Reduced Order Models
Highlights
[1] Biancolini, M. E., & Valentini, P. P. (2018). Virtual human bone modelling by interactive sculpting, mesh morphing and force-feedback. International Journal on Interactive Design and Manufacturing (IJIDeM), 12(4), 1223-1234.
[2] Capellini, K., Vignali, E., Costa, E., Gasparotti, E., Biancolini, M. E., Landini, L., ... & Celi, S. (2018). Computational fluid dynamic study for aTAA hemodynamics: an integrated image-based and radial basis functions mesh morphing approach. Journal of biomechanical engineering, 140(11).
[3] Biancolini, M. E., Ponzini, R., Antiga, L., & Morbiducci, U. (2012). A new workflow for patient specific image-based hemodynamics: parametric study of the carotid bifurcation. Computational Modelling of Objects Represented in Images III: Fundamentals, Methods and Applications, CRC Press, Rome, Italy.
Contacts:
Prof. Marco Evangelos Biancolini (biancolini@ing.uniroma2.it)
Dr. Corrado Groth (corrado.groth@uniroma2.it)
Dr. Ubaldo Cella (ubaldo.cella@uniroma2.it)
University of Rome “Tor Vergata”, Department of Enterprise Engineering “Mario Lucertini”
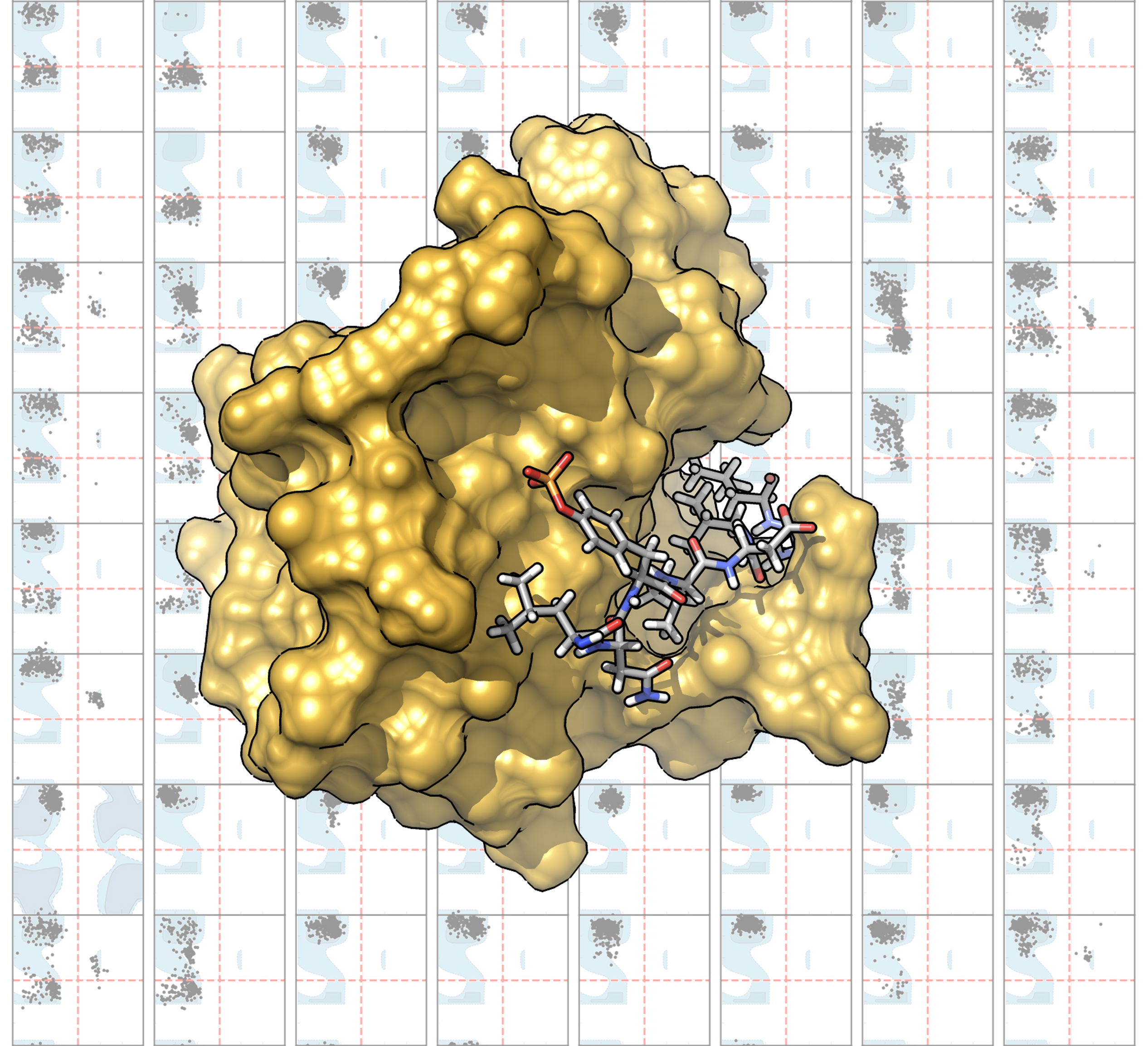
Structure of a complex between an octapeptide comprising a phosphotyrosine and the SH2 domain of the SHP2 protein from Molecular Dynamics simulations. On the background the Ramachandran Plots for different peptide sequences sampled during independent simulations.
Biophysical Chemistry
The group investigates complex systems of biological interest mainly by means of molecular dynamics simulations based on classical force fields, often compared with spectroscopic experiments. The main interests are: i) proteins involved in pathologies; ii) peptides and peptidomimetics in homogeneous and heterogeneous solutions; iii) nanosystems for biomedical applications; iv) modified carbohydrates for controlled drugs release.
Keywords
Molecular Dynamics Simulations, Proteins and Peptides, Polysaccharides, Nanosystems.
Highlights:
[1] Anselmi, M., Calligari, P., Hub, J., Tartaglia, M., Bocchinfuso, G., Stella, L., Structural Determinants of Phosphopeptide Binding to the N-Terminal Src Homology 2 Domain of the SHP2 Phosphatase. J. Chem. Inf. Mod. 2020 in press (https://doi.org/10.1021/acs.jcim.0c00307)
[2] Biscaglia, F., Ripani, G., Rajendran, S., Benna, C., Mocellin, S., Bocchinfuso, G., Meneghetti, M., Palleschi, A., Gobbo, M. Gold Nanoparticle Aggregates Functionalized with Cyclic RGD Peptides for Targeting and Imaging of Colorectal Cancer Cells. ACS Applied Nano Materials. 2019, 2:6436−6444
[3] Konai, M.M., Samaddar, S., Bocchinfuso, G., Santucci, V., Stella, L., Haldar, J. Selectively Targeting Bacteria through Tuning Molecular Design of Membrane-Active Peptidomimetic Amphiphiles. Chem Comm 2018, 54:4943.
Contacts:
Prof. Antonio Palleschi antonio.palleschi@uniroma2.it
Prof. Gianfranco Bocchinfuso bccgfr00@uniroma2.it
Prof. Lorenzo Stella stella@stc.uniroma2.it
Dr. Paolo Calligari paolo.calligari@uniroma2.it
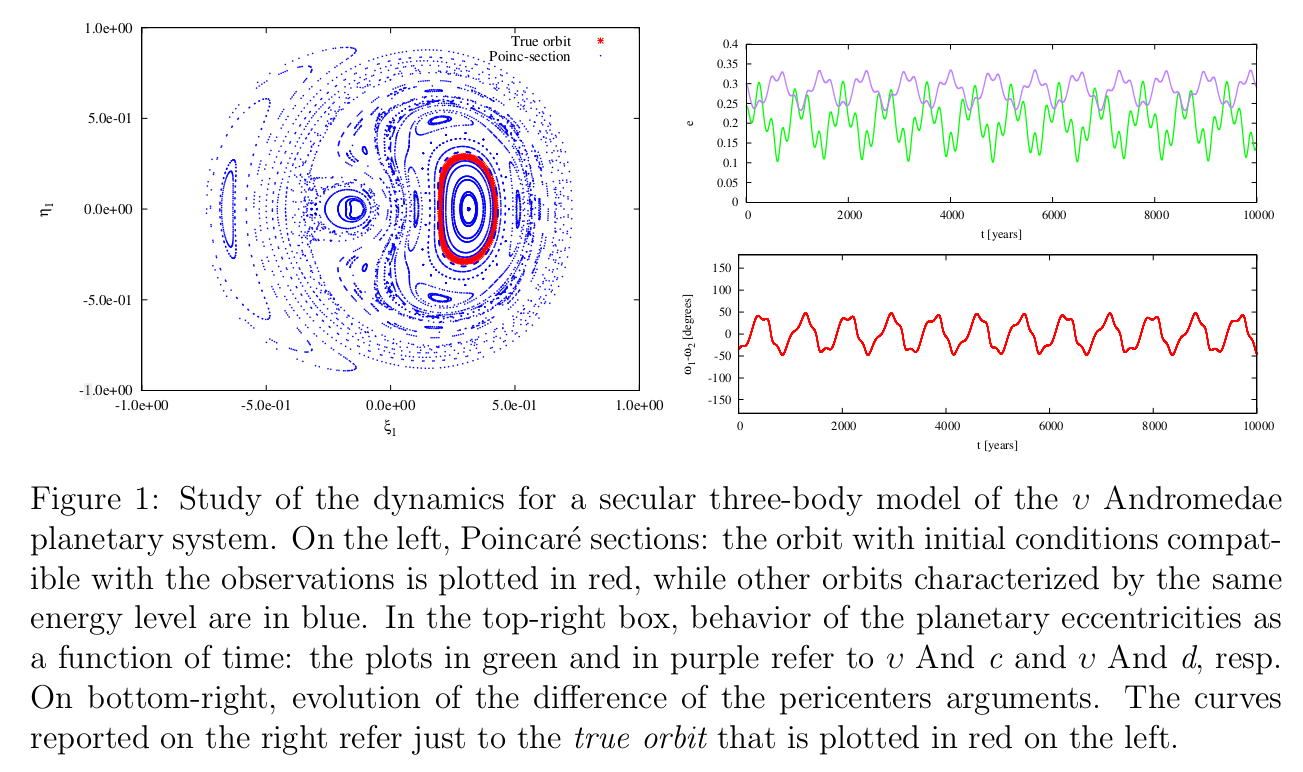
Celestial Mechanics
Nowadays, the research activity focusing on extrasolar systems that host more than one exoplanet is very exciting. These objects are currently studied into the framework of several fundamental sciences: Physics, Chemistry, Biology, etc. From an astronomical point of view, their characteristics are often very different with respect to those observed in our Solar System. There are several examples of exoplanets much more massive than Jupiter, orbiting close to the star and/or in quite eccentric orbits, some of them are in mean motion resonances, etc. Detection methods are presently unable to measure all the orbital elements, therefore the dynamical architecture of those systems is not yet completely understood and is currently under investigation. The usual strategy based on Hamiltonian perturbation theory can be successfully applied on reverse: we assume that those extrasolar planetary systems are long-term stable and we derive ranges of values for the unknown orbital elements and parameters (or, at least, we refine their knowledge), so that they are compatible with the prescribed stability. Such an approach can be efficiently implemented by explicitly constructing suitable Hamiltonian normal forms. This usually requires very relevant computational efforts and the use of algebraic manipulators that are developed ad hoc for this kind of problems.
Keywords
Stability of planetary systems, Hamiltonian perturbation theory, KAM theory, Computer Algebra Software.
Highlights:
[1] Giorgilli, A., Locatelli, U., Sansottera, M., "On the convergence of an algorithm constructing the normal form for lower dimensional elliptic tori in planetary systems", Cel. Mech. Dyn. Astr., 119, 397-424 (2014), https://doi.org/10.1007/s10569-014-9562-7
[2] Volpi, M., Locatelli, U., Sansottera, M., "A reverse KAM method to estimate unknown mutual inclinations in exoplanetary systems", Cel. Mech. Dyn. Astr., 130:36 (2018), https://doi.org/10.1007/s10569-018-9829-5
[3] Caracciolo, C., Locatelli, U., Sansottera, M., Volpi, M., "Librational KAM tori in the secular dynamics of the ʊ Andromedae planetary system", submitted (2021)
Contacts:
Prof. Ugo Locatelli locatell@mat.uniroma2.it
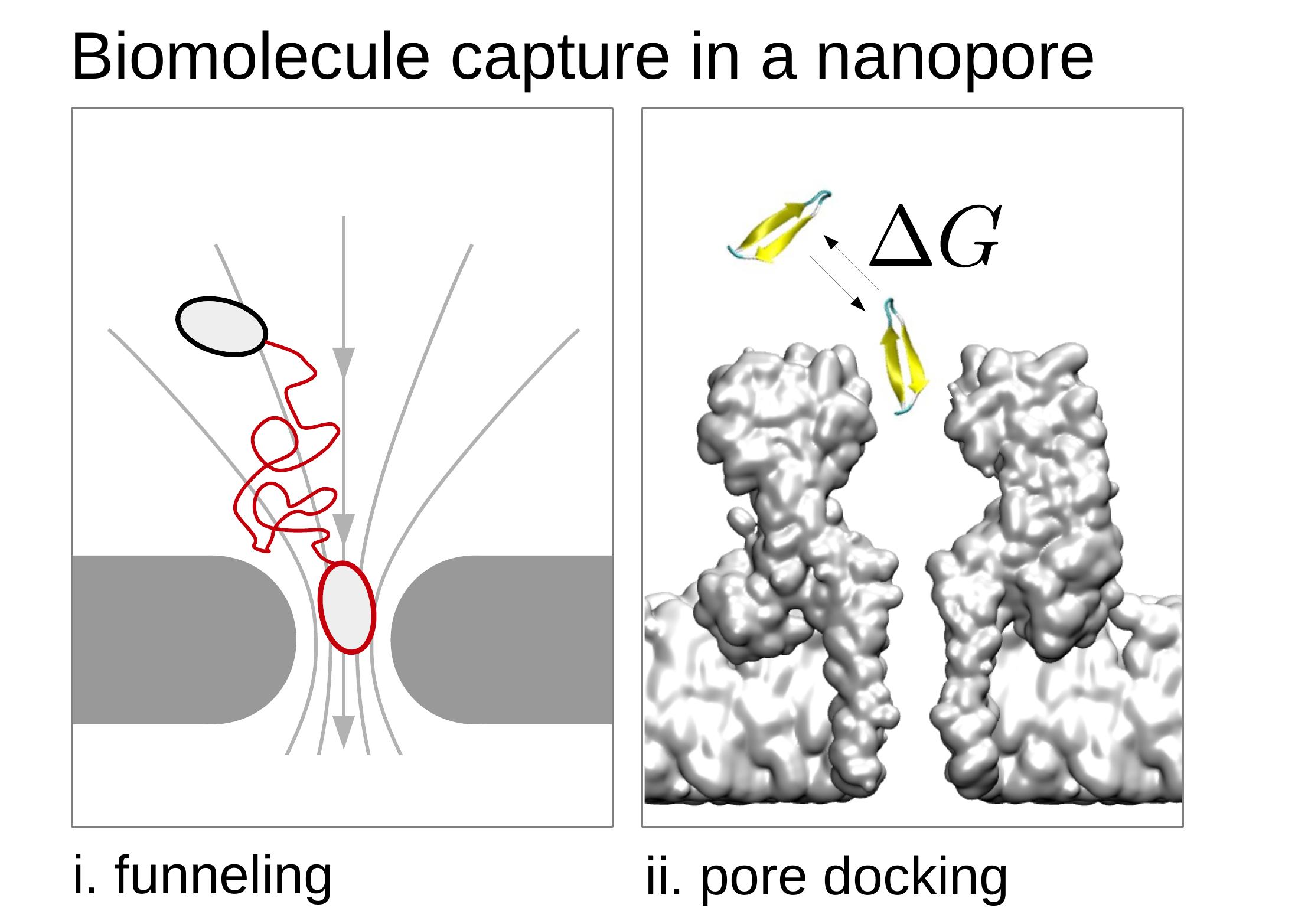
Far from the pore, the particle dynamics is ruled by the competition between diffusion and electric and hydrodynamic forces. When the particle reaches the pore entrance, pore-particle interactions become crucial and atomistic models are needed.
Nanopore and nanoporous membranes
Nanopore based devices have been revolutionizing several key-applications such as biosensing, energy harvesting and water treatment. Technological advancements are hindered by the lack of understanding and control of the coupling between hydrodynamics, electrokinetic and chemical effects that govern the flow through nanopores and the nanoparticle/biomolecule motion under nanoconfinement. We employ atomistic simulation, Brownian dynamics and continuum models to study the ion and solvent flow in biological and solid-state nanopores and the interaction between dispersed nanoparticles/biomolecules and the pore. The activity of the team is equally distributed between fundamental research (e.g. electrohydrodynamic coupling in nanopores) and applications (e.g. nanopore proteins sequencing, detection of toxic species in Li-S batteries)
Keywords: nanofluidics, biosensing, electroosmosis, blue-energy harvesting, atomistic simulation, coarse grained approaches, FEM.
Highlights:
[1] Di Muccio, G., della Rocca, B.M. and Chinappi, M., 2021. Geometrically Induced Selectivity and Unidirectional Electroosmosis in Uncharged Nanopores. arXiv preprint arXiv:2104.03390. https://arxiv.org/abs/2104.03390
[2] Chinappi, M., Yamaji, M., Kawano, R. and Cecconi, F., 2020. Analytical model for particle capture in nanopores elucidates competition among electrophoresis, electroosmosis, and dielectrophoresis. ACS nano, 14(11), pp.15816-15828.
https://pubs.acs.org/doi/abs/10.1021/acsnano.0c06981
[3] Bétermier, F., Cressiot, B., Di Muccio, G., Jarroux, N., Bacri, L., Della Rocca, B.M., Chinappi, M., Pelta, J. and Tarascon, J.M., 2020. Single-sulfur atom discrimination of polysulfides with a protein nanopore for improved batteries. Communications Materials, 1(1), pp.1-11. https://www.nature.com/articles/s43246-020-00056-4
[4] Bonome, E.L., Cecconi, F. and Chinappi, M., 2019. Translocation intermediates of ubiquitin through an α-hemolysin nanopore: implications for detection of post-translational modifications. Nanoscale, 11(20), pp.9920-9930.
https://pubs.rsc.org/en/content/articlelanding/2019/nr/c8nr10492a/unauth
Contacts:
Prof. Mauro Chinappi. mauro.chinappi@uniroma2.it
Dr. Blasco Morozzo della Rocca. blasco.morozzo.della.rocca@uniroma2.it
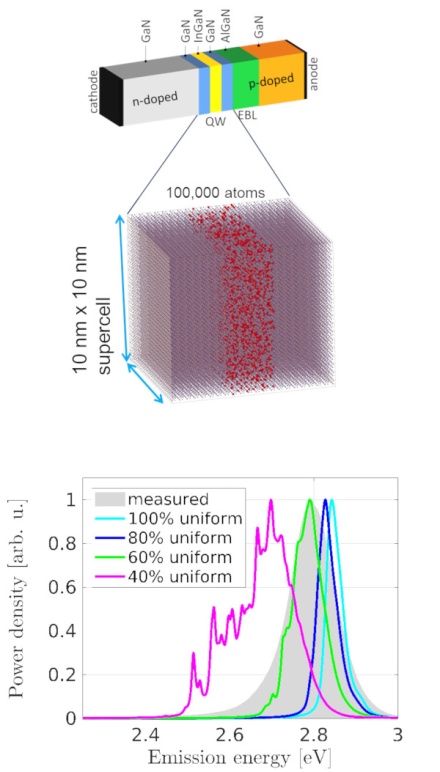
Multiscale/multiphysics simulation of optoelectronic devices
The Opto- and Nanoelectronics group of the Departement of Electronic Engineering has more than 15 years of experience in simulation and modelling of electronic devices, with special focus on III-nitride LEDs and HEMTs, organic LEDs and solar cells (dye sensitized, organic and hybrid perovskites). The group is developing a multiscale and multiphysics simulation software (tibercad) with the aim of integrating device level continuum models like linear elasticity, drift-diffusion or k·p with atomistic models like valence force field and empirical tight binding. The simulation framework has been applied during the last years to InGaN/GaN quantum-well LEDs, in order to study the effects of alloy disorder on device performance, and to perovskite based solar cell structures to simulate the optical properties of structures containing inclusions of different crystalline structure. In addition, the group is studying the light emission properties of micro- and nanoLEDs, using both full wave and ray-tracing approaches, with the aim of optimizing LED structures in terms of optical properties for specific application contexts, and part of the activity is devoted to ab-initio studies for perovskite based devices. The group runs a small HPC cluster with 12 CPU nodes and one GPU node (Nvidia Ampere A100).
Keywords: device simulation, mutiscale modelling, III-nitrides, photovoltaics, empirical tight-binding, alloy disorder
Highlights:
[1] A. Di Vito, A. Pecchia, A. Di Carlo, and M. Auf der Maur. Simulating random alloy effects in III-nitride light emitting diodes. Journal of Applied Physics, 128(4), 2020.
[2] A. Di Vito, A. Pecchia, M. Auf der Maur, and A. Di Carlo. Nonlinear work function tuning of lead-halide perovskites by Mxenes with mixed terminations. Advanced Functional Materials, 30, 1909028, 2020.
[3] A. Agresti, A. Pazniak, S. Pescetelli, A. Di Vito, D. Rossi, A. Pecchia, M. Auf der Maur, A. Liedl, R. Larciprete, Denis V. Kuznetsov, D. Saranin, and A. Di Carlo. Titanium-carbide Mxenes for work function and interface engineering in perovskite solar cells. Nature Materials, 18:1228–1234, 2019.
[4] M. Auf der Maur, A. Pecchia, G. Penazzi, W. Rodrigues, and A. Di Carlo. Efficiency drop in green InGaN/GaN light emitting diodes: The role of random alloy fluctuations. Physical Review Letters, 116(2), 2016.
Contacts:
Prof. Matthias Auf der Maur
Prof. Aldo Di Carlo